GenBio-PathFM: A State-of-the-Art Foundation Model for Histopathology
GenBio-PathFM: A State-of-the-Art Foundation Model for Histopathology
Kapse, S.; Aygün, M.; Cole, E.; Lundberg, E.; Song, L.; Xing, E. P.
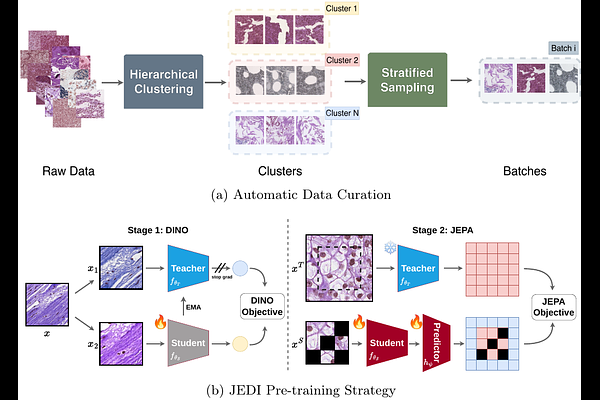
AbstractRecent advancements in histopathology foundation models (FMs) have largely been driven by scaling the training data, often utilizing massive proprietary datasets. However, the long-tailed distribution of morphological features in whole-slide images (WSIs) makes simple scaling inefficient, as common morphologies dominate the learning signal. We introduce GenBio-PathFM, a 1.1B-parameter FM that achieves state-of-the-art performance on public benchmarks while using a fraction of the training data required by current leading models. The efficiency of GenBio-PathFM is underpinned by two primary innovations: an automated data curation pipeline that prioritizes morphological diversity and a novel dual-stage learning strategy which we term JEDI (JEPA + DINO). Across the THUNDER, HEST, and PathoROB benchmarks, GenBio-PathFM demonstrates state-of-the-art accuracy and robustness. GenBio-PathFM is the strongest open-weight model to date and the only state-of-the-art model trained exclusively on public data.