methbiome: reference-based DNA methylation profiling of long-read metagenomic data
methbiome: reference-based DNA methylation profiling of long-read metagenomic data
Costeira, R.; Eustratiou Wong, E.; Bell, J. T.
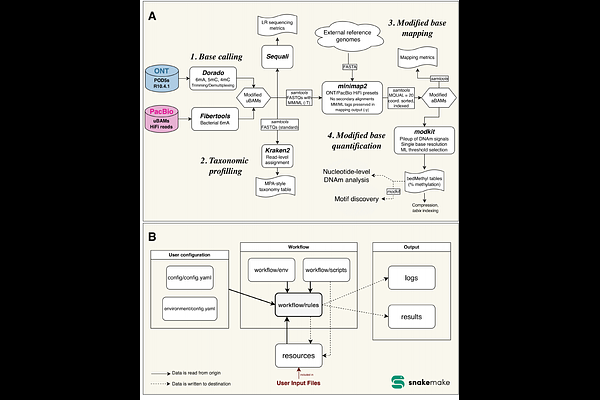
AbstractSummary: Long-read DNA sequencing technologies collect base modification information, but their capacity to profile epigenetic signals has not been widely explored within metagenomic context. Here, we present a novel pipeline consolidating key steps of meta-epigenomic profiling. methbiome can process both Oxford Nanopore and PacBio long-reads within a reference-based framework, facilitating cross-platform and between-sample comparison of DNA methylation signals in whole microbiomes. The tool is applicable to both microbial communities and isolates, for analyses within both methylation motifs as well as out-of-motif contexts, and can be used in metaepigenome-wide association studies. Availability and Implementation: methbiome was implemented using Snakemake and is available on GitHub (github.com/ricardocosteira/methbiome).