t2pmhc: A Structure-Informed Graph Neural Network to predict TCR-pMHC Binding
t2pmhc: A Structure-Informed Graph Neural Network to predict TCR-pMHC Binding
Polster, M.; Stadelmaier, J.; Ball, E.; Scheid, J.; Bauer, J.; Nelde, A.; Claassen, M.; Dubbelaar, M. L.; Walz, J. S.; Nahnsen, S.
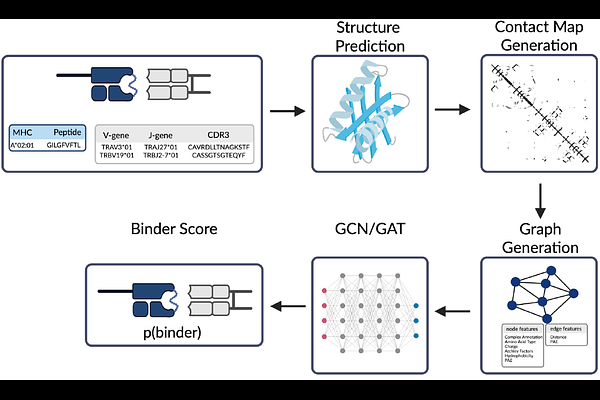
AbstractMapping of T cell receptors (TCRs) to their cognate MHC-presented peptides (pMHC) is central for the development of precision immunotherapies and vaccine design. However, accurate prediction of TCR affinity to peptide antigens remains an open challenge. Most approaches rely solely on sequence information, although increasing evidence suggests that TCR-pMHC binding is primarily determined by three-dimensional structural interactions within the entire TCR-pMHC complex. Consequently, sequence-based methods often fail to generalize to peptides not included in the training data (unseen peptides). Here we introduce t2pmhc, a structure-based graph neural network framework for predicting TCR-pMHC binding using predicted structures of the entire TCR-pMHC complex. We evaluated a Graph Convolutional Network (GCN) and a Graph Attention Network, both demonstrating improved generalization to unseen peptides compared to state-of-the-art models across a variety of public datasets. Evaluation with crystallographic structures yields high-confidence predictions, indicating that current limitations of structure-based models are largely driven by the accuracy of structure prediction. Analysis of node attention patterns in t2pmhc-GCN reveals biologically consistent patterns, assigning high attention to the peptide and the CDR3 regions. Within the peptide sequence, canonical MHC anchor residues are consistently downweighted, whereas potential TCR-binding residues are upweighted. These findings establish t2pmhc as a structure-informed framework for robust TCR-pMHC binding prediction, enabling improved generalization to unseen antigens and providing a foundation for integrating TCR repertoire sequencing into vaccine design and immunotherapy.