MERFISH 2.0, an ultra-sensitive single-cell spatial transcriptomics imaging chemistry across diverse tissue types
MERFISH 2.0, an ultra-sensitive single-cell spatial transcriptomics imaging chemistry across diverse tissue types
He, J.; He, L.; Wang, B.; Wiggin, T.; Chen, R.; Wang, H.; Yang, B.; Tattikota, S. G.; Maziashvili, L.; Zhang, T.; Revuru, S.; Wang, S.; Patil, S.; Sun, Y.; Sun, Y.; Li, M.; Cai, Y.; Wu, L.; Pentrenko, N.; Vasaturo, A.; Ray, M.; Emanuel, G.
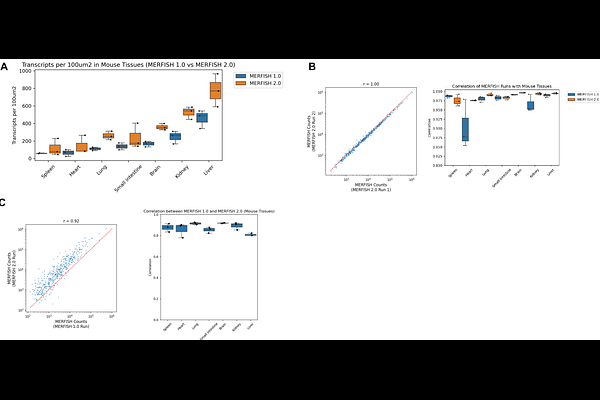
AbstractSpatial transcriptomics has emerged as a transformative approach for elucidating tissue architecture, cellular heterogeneity, and disease mechanisms by preserving the spatial context of gene expression in cells. Despite these advances, many spatial transcriptomic methods underperform in archival or clinically relevant specimens, particularly formalin-fixed, paraffin-embedded (FFPE) tissues, where RNA degradation and crosslinking hinder transcript detection. To address these challenges, we developed Multiplexed Error Robust Fluorescence In Situ Hybridization 2.0 (MERFISH 2.0), an optimized spatial transcriptomic imaging chemistry to enhance profiling of fragmented and highly crosslinked RNA. Across diverse human and mouse tissues preserved as fresh-frozen, fixed-frozen, and FFPE specimens, MERFISH 2.0 substantially increased transcript detection sensitivity by up to ~8-fold relative to MERFISH 1.0 while preserving quantitative concordance. In archived fresh frozen human brain samples, MERFISH 2.0's enhanced sensitivity improved transcript recovery, enhanced cell type resolution and spatial analyses. In low quality archival FFPE human breast cancer specimen, MERFISH 2.0 revealed additional cell populations, novel cell clusters, refined tumor-immune architecture, and increased detection of gene-gene and cell-cell interactions relative to MERFISH 1.0, underscoring the impact of improved sensitivity on downstream spatial analysis. By substantially expanding robust transcript detection to degraded and archival samples, MERFISH 2.0 enables scalable, cohort-level spatial transcriptomic analysis across clinically relevant tissue collections.