G-VEP: GPU-Accelerated Variant Effect Prediction for Clinical Whole-Genome Sequencing Analysis
G-VEP: GPU-Accelerated Variant Effect Prediction for Clinical Whole-Genome Sequencing Analysis
Green, E.; Mardinoglu, A.
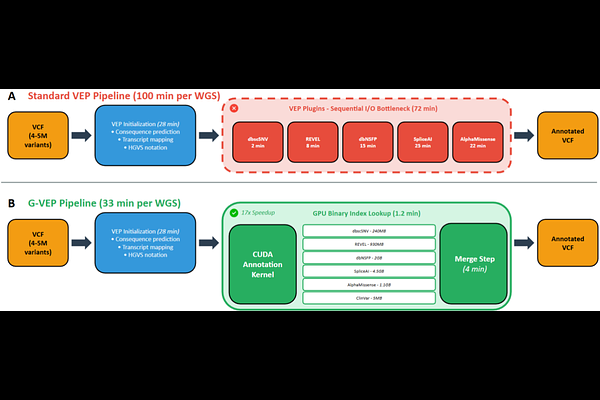
AbstractWhole-genome sequencing (WGS) has transformed clinical diagnostics, yet variant annotation remains a computational bottleneck. The Variant Effect Predictor (VEP) integrates pathogenicity predictors and population databases essential for ACMG/AMP variant classification, but these annotation plugins are fundamentally I/O-bound, consuming over 70% of total pipeline runtime. Here, we present G-VEP, a GPU-accelerated annotation framework built on a custom CUDA kernel that replaces sequential per-variant database lookups with massively parallel binary search across precomputed indices. By executing annotation lookups for all input variants simultaneously, G-VEP reduces plugin runtime from 72 minutes to 4 minutes (17-fold acceleration) and total annotation runtime from 100 minutes to 33 minutes (3-fold acceleration), while maintaining complete concordance with standard VEP output. Benchmarking across 75 clinical WGS samples demonstrated consistent performance, with no annotation discrepancies; validation on samples containing known pathogenic variants confirmed the preservation of all clinically significant findings. The 8.8 GB index footprint fits within consumer-grade 16 GB GPUs. G-VEP addresses an unmet need in clinical WGS analysis, while GPU suites such as NVIDIA Parabricks accelerate alignment and variant calling, they do not provide the Ensembl VEP plugin ecosystem used in clinical interpretation. G-VEP removes this final bottleneck and enables accelerated WGS interpretation. G-VEP is freely available through a web-based user interface with REST API documentation at https://www.phenomeportal.org/gvep, and source code for local installation and deployment at https://github.com/Phenome-Longevity/G-VEP.