Rhizobacterial Biosensors Spatially Map Natural and Engineered Sucrose Exudation
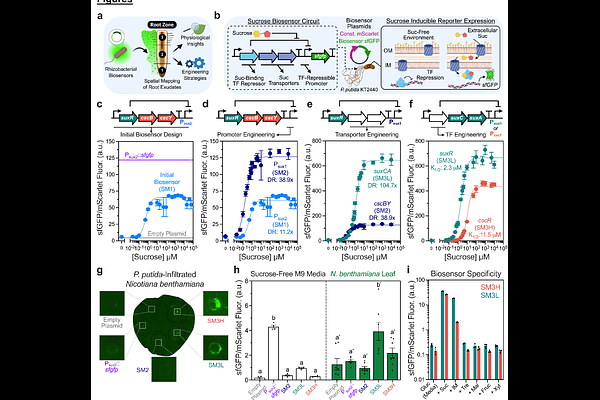
Rhizobacterial Biosensors Spatially Map Natural and Engineered Sucrose Exudation
Dundas, C. M.; Brinkman, G. A.; Clarke, T.; Payne, M.; Ureta, J. A.; Velasco, I.; Wallace, J. G.; Dinneny, J. R.
AbstractRoot exudation mediates the delivery of plant primary and secondary metabolites into soil, where they regulate plant-microbe interactions and terrestrial carbon cycling. Conventional exudate analyses quantify total root-released carbon, yet obscure the spatial origin and rhizosphere influence of individual compounds. Here, we develop a rhizobacterial biosensor platform, named Suc-MAPP, to map local exudate profiles along the surface of colonized root tissues. Focusing on sucrose, we engineered sfGFP-based, sucrose-responsive gene circuits in Pseudomonas putida KT2440 for live imaging of exudate concentrations in the micromolar range. These biosensors reveal spatially structured sucrose exudation patterns across eudicots and monocots and implicate photoassimilate source-sink dynamics as a major determinant. We further apply this platform to phenotype exudation modulated by synthetic gene circuitry in Arabidopsis thaliana, identifying genetic design rules for graded sucrose release and quantifying how engineered export sculpts rhizosphere assembly of a defined bacterial community. Together, these results establish programmable rhizobacterial biosensors as tools to spatially resolve plant-environment carbon exchange in situ and provide a framework for extending this approach to diverse exudate targets.