DNA fragment length analysis using machine learning assisted vibrational spectroscopy
DNA fragment length analysis using machine learning assisted vibrational spectroscopy
Fatayer, R.; Ahmed, W.; Szeto, I.; Sammut, S.-J.; Senthil Murugan, G.
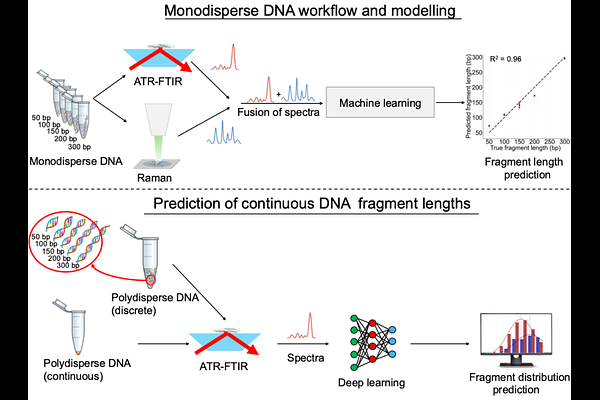
AbstractDNA length analysis is essential for genomic workflows including next-generation sequencing and fragmentomics based diagnostics. Conventional approaches typically require large, expensive instrumentation and sample-destructive protocols with long processing times. Here we present a rapid, label-free approach integrating vibrational spectroscopy with deep learning to quantify DNA fragment length distributions. We demonstrate that ATR-FTIR and Raman spectroscopy capture length-dependent spectral features arising from phosphate backbone, nucleobase, and structural vibrations. Machine learning models trained on spectra acquired from purified monodisperse DNA (50-300 bp) predicted DNA length with high accuracy (R2=0.92-0.94), with multimodal fusion improving performance to R2=0.96. A convolutional neural network trained on 35 DNA mixtures comprising molecules of different lengths also successfully deconvoluted their fragment length profile. Transfer learning enabled adaptation to biological samples, achieving low prediction error (RMSE=0.3-7.2%, {Delta}=12 bp). Importantly, the method requires only 4 L sample and 15 minutes passive drying, with no consumables beyond cleaning materials, and allows full sample recovery. This establishes vibrational spectroscopy as a scalable alternative for DNA length quantification.