Pareto optimization of masked superstrings improves compression of pan-genome k-mer sets
Pareto optimization of masked superstrings improves compression of pan-genome k-mer sets
Plachy, J.; Sladky, O.; Brinda, K.; Vesely, P.
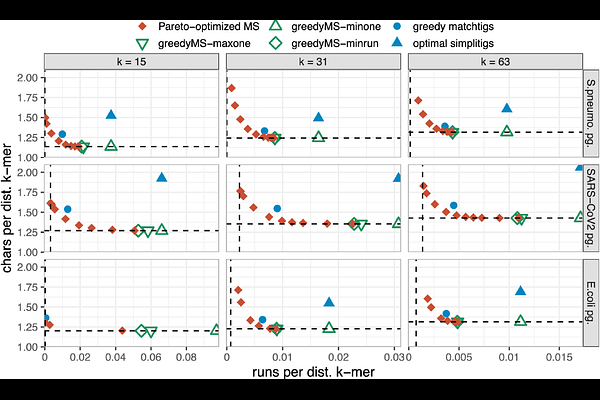
AbstractThe growing interest in k-mer-based methods across bioinformatics calls for compact k-mer set representations that can be optimized for specific downstream applications. Recently, masked superstrings have provided such flexibility by moving beyond de Bruijn graph paths to general k-mer superstrings equipped with a binary mask, thereby subsuming Spectrum-Preserving String Sets and achieving compactness on arbitrary k-mer sets. However, existing methods optimize superstring length and mask properties in two separate steps, possibly missing solutions where a small increase in superstring length yields a substantial reduction in mask complexity. Here, we introduce the first method for Pareto optimization of k-mer superstrings and masks, and apply it to the problem of compressing pan-genome k-mer sets. We model the compressibility of masked superstrings using an objective that combines superstring length and the number of runs in the mask. We prove that the resulting optimization problem is NP-hard and develop a heuristic based on iterative deepening search in the Aho-Corasick automaton. Using microbial pan-genome datasets, we characterize the Pareto front in the superstring-length/mask-run space and show that the front contains points that Pareto-dominate simplitigs and matchtigs, while nearly encompassing the previously studied greedy masked superstrings. Finally, we demonstrate that Pareto-optimized masked superstrings improve pan-genome k-mer set compressibility by 12-19% when combined with neural-network compressors.