A Multi-Dataset Transcriptomic Analysis Unravels Core Mechanisms Involving Vitamin D Metabolism and Inflammatory Pathways for Frailty Diagnosis.
A Multi-Dataset Transcriptomic Analysis Unravels Core Mechanisms Involving Vitamin D Metabolism and Inflammatory Pathways for Frailty Diagnosis.
Hu, X.; Zheng, W.; Li, Y.; Zhou, D.
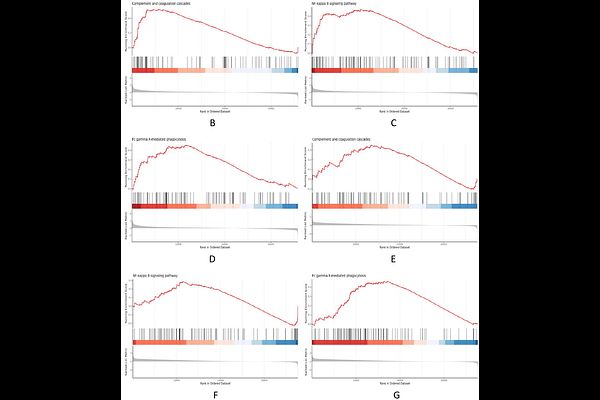
AbstractFrailty is a prevalent geriatric syndrome, and the shortage of objective biomarkers restricts its early diagnosis and intervention. This study aimed to identify robust molecular signatures and diagnostic markers for frailty using bioinformatics analyses of multiple independent datasets. Two transcriptome datasets (GSE144304, n=80; GSE287726, n=70) were obtained from the GEO database. We performed differential gene expression analysis, GO, KEGG and GSEA enrichment, and machine learning (70% training / 30% validation) to screen and validate core biomarkers. Numerous shared differentially expressed genes were identified. Vitamin D metabolism, ABC transporter, and inflammatory/immune pathways were consistently enriched and confirmed by GSEA. Machine learning models based on these signatures showed favorable diagnostic performance. Our study demonstrates that vitamin D metabolic disorders and chronic inflammation are core molecular features of frailty. The identified biomarkers provide new strategies for basic research, early clinical diagnosis, and therapeutic target development for frailty.