FourC: identifying significant and differential contacts in 1D chromatin conformation data
FourC: identifying significant and differential contacts in 1D chromatin conformation data
Wong, W.; Kaplan, S. J.; Luo, R.; Pulecio Rojas, J. A.; Yan, J.; Huangfu, D.; Leslie, C. S.
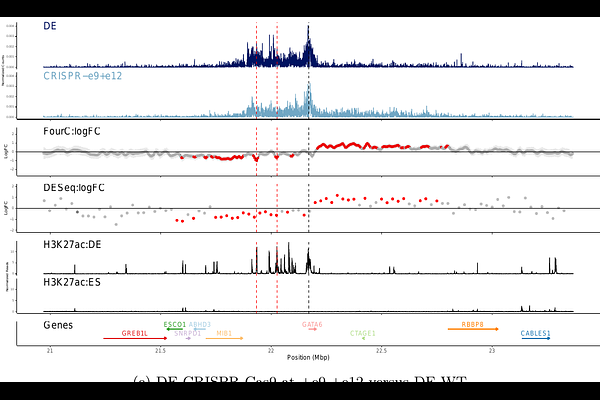
Abstract4C-seq is a cost-effective 3C-based assay that measures the interactions between a single genomic element and all other genomic elements. However, 4C-seq data remains semi-quantitative because it cannot be deduplicated without UMIs. To address this, we developed an open source method, FourC, based on a Bayesian Bernoulli regression model, that overcomes the duplication problem and models spatial patterns with Gaussian processes to identify significantly enriched and differential contacts. We demonstrate the utility of FourC on 4C-seq data that profiles the local chromatin structure at key genes necessary for pancreatic differentiation and under CRISPR perturbation of enhancers.