ExoFILT: Transfer learning for robust and accelerated analysis of exocytosis single-particle tracking data
ExoFILT: Transfer learning for robust and accelerated analysis of exocytosis single-particle tracking data
Kramer, E.; Betancur, L. I.; Meek, S.; Tosi, S.; Manzo, C.; Oliva, B.; Gallego, O.
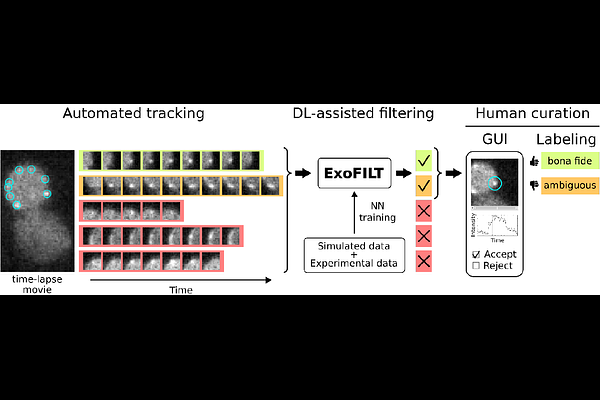
AbstractMotivation: Understanding constitutive exocytosis at the molecular level requires quantitative characterization of protein dynamics during the process. Single-particle tracking allows the measurement of protein dynamics in living cells. However, identifying bona fide exocytic events requires extensive manual annotation, limiting throughput and introducing personal biases that affect reproducibility. Results: We present ExoFILT, a deep learning-based classifier designed to identify exocytic events in single-particle tracking data, using the exocyst complex as a reference. Trained via transfer learning on simulated and experimental data, ExoFILT reduces the time required for manual annotation by ten-fold while improving measurement consistency across researchers. When applied to simultaneous dual-color time-lapse movies, ExoFILT enabled the systematic quantification of temporal relationships between exocytic proteins. The increased throughput uncovered distinct subpopulations of exocytic events with differential molecular composition (e.g., events with and without detectable levels of Sec1), underscoring the potential of ExoFILT to reveal mechanistic insights into exocytosis.