Shape2Fate: a morphology-aware deep learning framework for tracking endocytic and exocytic carriers at nanoscale.
Shape2Fate: a morphology-aware deep learning framework for tracking endocytic and exocytic carriers at nanoscale.
Harmanec, A.; Dagg, A. D.; Kamenicky, J.; Kerepecky, T.; Makieieva, Y.; Pereira, C.; Bright, N.; Menon, D.; Gershlick, D. C.; Vaskovicova, N.; Lai, T.; Fazakerley, D. J.; Schermelleh, L.; Sroubek, F.; Kadlecova, Z.
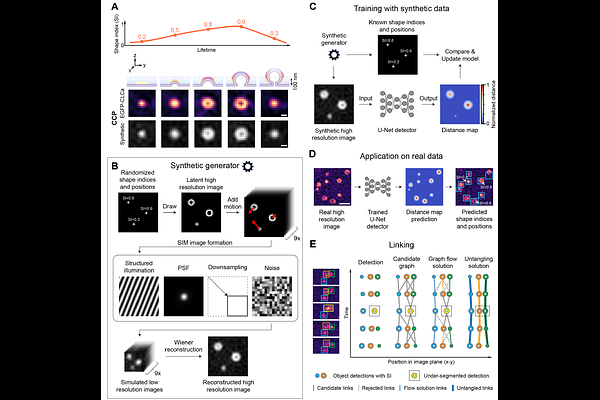
AbstractPlasma membrane homeostasis requires balanced exocytosis and endocytosis, yet their coordination at the single-event level in non-neuronal cells is unresolved. We present Shape2Fate, a morphology-aware deep-learning pipeline that detects, tracks, and classifies individual exocytic and endocytic carriers in live-cell total internal reflection fluorescence structured illumination microscopy (TIRF-SIM) movies at ~100 nm resolution. Trained on synthetic data and exploiting carrier shape evolution rather than fluorescence intensity, Shape2Fate achieves expert-level tracking and outcome classification across diverse cell types, imaging conditions, and microscope platforms. Applying Shape2Fate to constitutive secretion and insulin-stimulated GLUT4 exocytosis in adipocytes, we uncover two opposing exo-endocytic coupling architectures: exocytic fusion locally nucleates de novo clathrin-coated pits, whereas GLUT4 vesicles target pre-existing pits for rapid cargo capture. These findings establish that the spatial rules governing exo-endocytic coordination are not universal but are pathway-specific. Shape2Fate is openly available, enabling direct event-level mechanistic dissection of exo-endocytic coordination across pathways in living cells.