Cross-Platform Transcriptomic Validation Identifies SERPINB2 as a Robust Chondrogenic Biomarker and Reveals Coordinated SERPIN Network Activation During Cartilage Lineage Commitment
Cross-Platform Transcriptomic Validation Identifies SERPINB2 as a Robust Chondrogenic Biomarker and Reveals Coordinated SERPIN Network Activation During Cartilage Lineage Commitment
Gonzalez-Reyes, B. E.; Hernandez-Lopez, E.; Leyva-Gonzalez, G.; Herrera-Camarena, M. C.; Gonzalez-Ruiz, A. G.; Pena-Rodriguez, L. L.; Espinosa-Morales, C.; Rojas-Berges, I.; Villamil-Galvan, R. M.; Estrada-Elorza, M. d. C.; Martinez-Nava, G. A.; Martinez-Mayorga, K.; Cuz-Lemini, M.; Granados-Montiel, J.
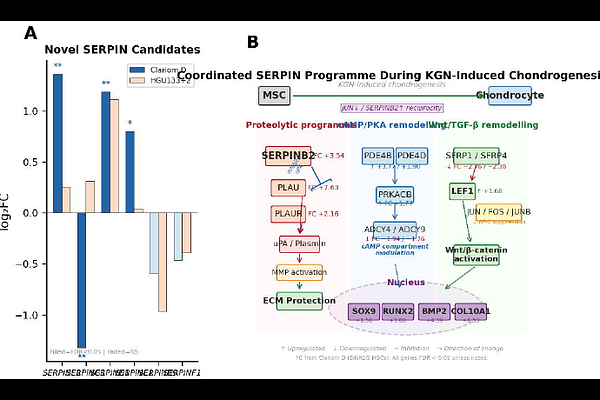
AbstractObjective To validate SERPINB2 and SERPINA9 as chondrogenic biomarker candidates across independent transcriptomic platforms and cell sources, to characterise the complete SERPIN expression landscape during kartogenin (KGN)-induced chondrogenic differentiation of human mesenchymal stem cells (hMSCs), and to identify novel SERPIN biomarker candidates and their signalling context during cartilage lineage commitment. Design Multi-platform transcriptomic analysis across three independent datasets: (i) Affymetrix HGU133+2 microarray of KGN-induced chondrocytes versus undifferentiated hMSCs (ATCC source); (ii) Affymetrix Clariom D whole-transcriptome array of KGN-treated versus control hMSCs from an independent Mexican source (SINREG Laboratories); and (iii) previously published qPCR validation. Differential expression was computed using limma with Benjamini,Hochberg correction. SERPIN-focused cross-platform correlation and targeted pathway analysis were performed. Results The Clariom D dataset yielded 1,869 differentially expressed genes (925 upregulated, 944 downregulated; FDR < 0.05) from 29,124 transcripts tested. SERPINB2 was concordantly upregulated across all three platforms (Clariom D: fold-change [FC] +3.54, FDR = 0.006; HGU133+2: log2FC = +3.29, nominal P = 0.027; qPCR confirmed), establishing it as one of the most reproducible transcriptomic signals in chondrogenic differentiation. In the direct Bone versus Cart comparison, SERPINB2 showed ~45-fold chondrogenic enrichment (log2FC = -5.45, adjusted P < 0.0001). Cross-platform SERPIN correlation was significant (Pearson r = 0.54, P = 0.0025; n = 29 shared genes). Four additional SERPINs reached genome-wide significance on Clariom D: SERPINE2 (FC +2.57), SERPING1, SERPIND1, and SERPINE1. SERPINA9 was not replicated in the independent SINREG source, identifying it as a context-dependent marker. Conclusions SERPINB2 is a robust, cross-platform chondrogenic biomarker with translational potential for osteoarthritis (OA) monitoring. The coordinated SERPIN programme activates a multi-layered proteolytic and signalling network during cartilage lineage commitment, positioning SERPINB2 as a functional regulator of the chondro-osteogenic lineage decision.