Structural insights into target detection by the S. marcescens type III CRISPR complex and its deployment inSNP identification
Structural insights into target detection by the S. marcescens type III CRISPR complex and its deployment inSNP identification
Perdigao, C. C.; Ajisafe, L. O.; Sunny, A. T.; Wu, S.; Dokland, T.; Dunkle, J. A.
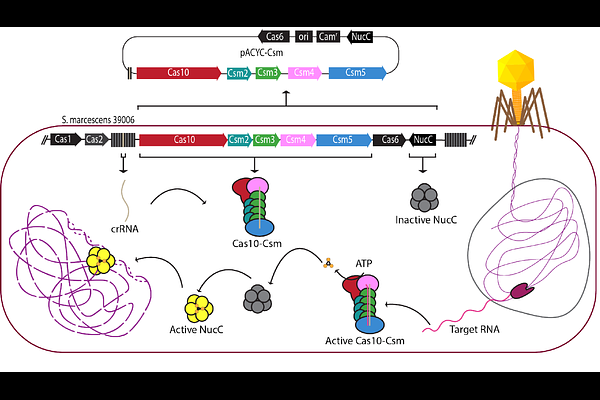
AbstractType III CRISPR systems utilize a complex containing Cas10, additional Cas proteins and a crRNA to detect foreign transcripts. Upon detection, Cas10 synthesizes cyclic oligoadenylates (cOA), signaling molecules that coordinate interference by stimulating downstream enzymes with DNase, RNase, protease or other activities. Type III systems are among the most abundant CRISPR systems in prokaryotes and understanding the structure-function relationships that control transcript detection and cOA synthesis will advance the understanding of the broader physiological roles of these systems. Type III systems possess properties well-suited to their deployment as molecule diagnostics: specific detection and activation of a cascade of multi-turnover enzymatic reactions that can be harnessed for signal generation. We determined that Serratia marcescens Cas10-Csm (SmCas10-Csm) synthesizes predominantly cA3 molecules and this synthesis is sensitive to mismatches in the crRNA-target RNA duplex adjacent to Cas10. We determined the structure of SmCas10 Csm unbound and bound to target RNA identifying conformational changes associated with target binding. We demonstrate that SmCas10-Csm can distinguish between single nucleotide polymorphisms that occur in the human HBB transcript that are associated with sickle cell disease indicating an additional role for type III CRISPR systems in point-of-care diagnostics in low-resource settings.