Decoding TF-Specific Predictability in Cross-Species Binding Site Inference
Decoding TF-Specific Predictability in Cross-Species Binding Site Inference
Wang, Y.; Liu, G.; Wang, Y.; Zhang, Y.
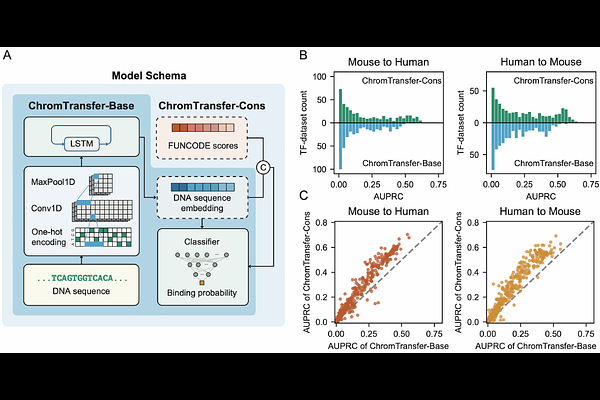
AbstractAccurately identifying transcription factor (TF) binding sites across species is essential for understanding conserved gene regulatory mechanisms. While experimental techniques such as ChIP-seq have enabled genome-wide TF-binding maps, their application is often constrained by the limited availability of high-quality antibodies. Computational approaches that leverage data from one species to predict TF-binding sites in other species have emerged as valuable alternatives. However, existing models often rely on uniform modeling assumptions, overlooking substantial variability in cross-species predictability across TFs. In this study, we systematically evaluated the cross-species predictability of 137 TFs using 425 human-mouse ChIP-seq dataset pairs matched by cell type, and identified key biological features underlying this variability. Building on these insights, we developed ChromTransfer, a TF-aware cross-species prediction framework that integrates DNA sequence, functional conservation, TF-specific co-binding signals, and shared chromatin context signals. These regulatory signals substantially improve prediction performance, particularly for TFs with weak or absent motif enrichment. Together, this study establishes a biologically informed and scalable framework for TF-specific cross-species TF-binding site prediction and provides a practical strategy for extending regulatory annotations across species.