ProteinSage: From implicit learning to explicit structural constraints for efficient protein language modeling
ProteinSage: From implicit learning to explicit structural constraints for efficient protein language modeling
Shen, L.; Chao, L.; Liu, T.; Liu, Q.; Zhou, G.; Wang, H.; Dong, X.; Li, T.; Zhang, X.; Ni, J.
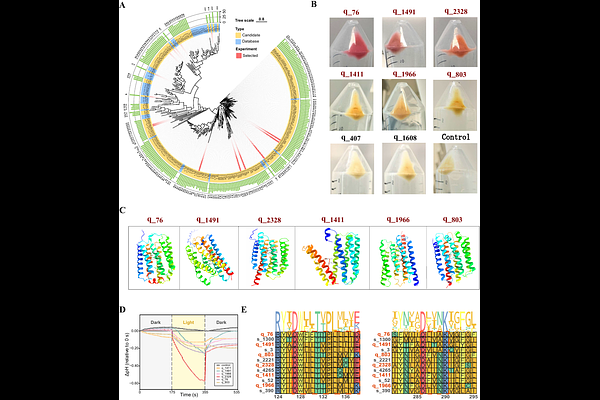
AbstractWhile protein language models typically rely on sequence-only pretraining objectives, this approach often fails to capture structural regularities and demands large datasets. To address this, we introduce ProteinSage, a pretraining framework that learns protein representations under explicit structural constraints. ProteinSage incorporates structural signals via structure-guided masking and a causal objective designed to model longrange dependencies. This structure-constrained pretraining equips ProteinSage with transferable representations using less data and computation, yet achieves competitive or superior performance across diverse structure-aware and general protein modeling benchmarks. To determine whether these gains stem from genuine structural generalization rather than task-specific fitting, we applied ProteinSage to a structure-driven protein discovery task, focusing on proteins with multi-pass transmembrane helical architectures such as distantly related microbial rhodopsins. The model successfully identified six previously unannotated microbial rhodopsin homologs. Together, our work establishes structure-constrained pretraining as an effective pathway toward data-efficient and structurally faithful protein representation learning.