Activity-based profiling of bacterial RNA-modifying enzymes reveals species-specific 5-methyluridine modification in Bacillus subtilis 23S rRNA
Activity-based profiling of bacterial RNA-modifying enzymes reveals species-specific 5-methyluridine modification in Bacillus subtilis 23S rRNA
Jaber, Q. Z.; Yu, N. J.; Masuda, I.; Hou, Y.-M.; Kleiner, R. E.
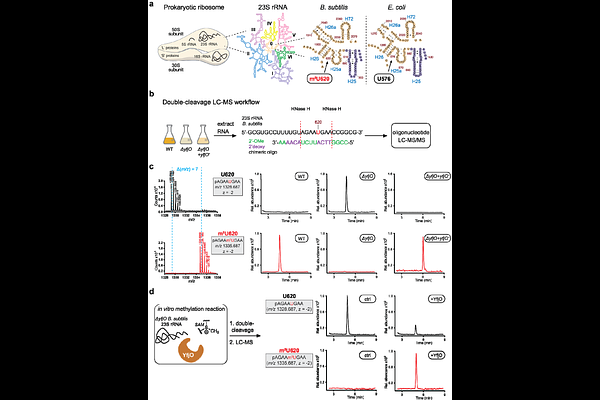
AbstractRNA modifications play an important role in biological processes. Mapping the diversity of RNA chemistry and studying the biological function of individual modifications remains an outstanding challenge in many organisms. In particular, RNA modifications remain poorly studied across most bacterial systems. Our group previously developed RNA-mediated activity-based protein profiling (RNABPP), a reactivity-based strategy employing metabolic labeling and quantitative proteomics to profile RNA modification writer enzymes in human cells. Here we adapt this approach to characterize RNA-modifying enzymes in bacteria. We apply metabolic labeling with 5-fluoropyrimidine nucleosides and phase separation-based enrichment of RNA-protein complexes (RNABPP-PS) to profile RNA pyrimidine modifying enzymes in E. coli and B. subtilis. We identify known and putative bacterial pyrimidine C5 methyltransferases, pseudouridine synthases, and dihydrouridine synthases, demonstrating the utility of our approach. Further, we find the carboxymethylaminomethyluridine (cnmn5U)-forming enzyme MnmG (GidA), supporting the existence of a covalent protein-RNA intermediate during the catalytic cycle. Finally, we use RNABPP-PS in B. subtilis to identify YfjO, an uncharacterized protein that is homologous to 5-methyluridine (m5U) methyltransferases. We use nucleoside and oligonucleotide mass spectrometry to establish that YfjO (which we rename as RlmS) installs m5U620 in the 23S rRNA (U576 in E. coli), a modification specific to the B. subtilis ribosome. We characterize {Delta}yfjO B. subtilis which has impaired cell proliferation and protein translation rate compared to WT. Taken together, our study establishes a versatile platform for RNA modifying enzyme discovery and characterization in bacteria and illuminates species-specific rRNA modification chemistry in B. subtilis.