A single-cell and spatial atlas of prostate cancer reveals the combinatorial nature of gene modules underlying lineage plasticity and metastasis
A single-cell and spatial atlas of prostate cancer reveals the combinatorial nature of gene modules underlying lineage plasticity and metastasis
Song, H.; Xu, J.; Velazquez-Arcelay, K.; Demirci, A.; Raizenne, B. L.; Hsu, S. C.; Choi, J.; Pham, J. H.; Chen, Y.-A.; Weinstein, H. N. W.; Salzman, I.; Tsui, M.; Akutagawa, J.; Adingo, W.; Goldschmidt, E.; Carroll, P. R.; Hong, J. C.; Heaphy, C. M.; Cooperberg, M. R.; Greenland, N.; Campbell, J. D.; Huang, F. W.
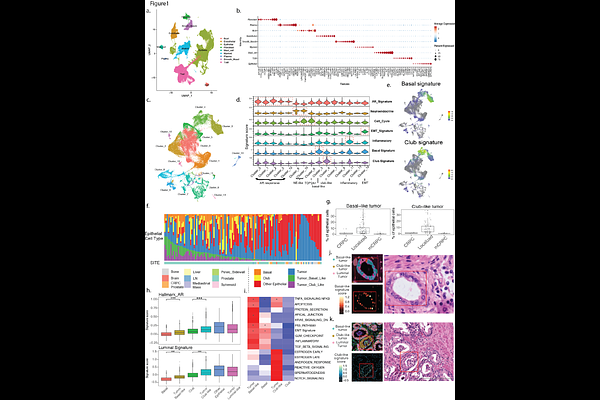
AbstractProstate cancer encompasses a spectrum of disease states driven by complex cellular heterogeneity. To delineate the transcriptional programs underlying lineage plasticity and metastasis, we constructed a comprehensive single-cell atlas of 128 patients, spanning localized, castration-resistant, and metastatic disease. Lineage plasticity was prevalent in localized disease, with subsets of tumor cells adopting distinct basal-like and club-like states. Luminal-like cancer cells also displayed extensive lineage infidelity, defined not by a binary loss of identity but by the combinatorial erosion of luminal gene modules associated with higher grade and stage. In the metastatic setting, gene program association analysis (GPAS) identified a broad induction of cell-cycle gene modules across organ sites as well as an induction of organ-specific gene modules, including osteomimetic signaling in bone, neuro-migratory genes in brain, and erythroid-like transitions in liver. Neuroendocrine prostate cancers (NEPCs) were not monolithic but defined by combinations of NE-associated gene modules including a novel HES6 program. Notably, these modules were detected at intermediate levels in localized samples, suggesting molecular plasticity precedes histological transformation. We also developed a refined NE signature that could distinguish NEPC tumors more accurately than previously published signatures. Within the tumor microenvironment (TME), we observed an elevation of pro-inflammatory Th17 T-cells in African American patients and identified a rare Schwann cell population. Finally, we present PCformer, a transformer-based foundation model trained on >500,000 cells to automate cell-state classification. Together, this comprehensive atlas demonstrates the complex nature of gene modules underlying lineage infidelity and plasticity in cancer cells and highlights distinct immune and stromal populations within the tumor ecosystem.