Bound or unbound: Mapping and monitoring receptor oligomerization using time-resolved fluorescence
Bound or unbound: Mapping and monitoring receptor oligomerization using time-resolved fluorescence
Greife, A.; Liu, R.; Koehler, P. S.; Heinze, K. G.; Hemmen, K.; Peulen, T.-O.
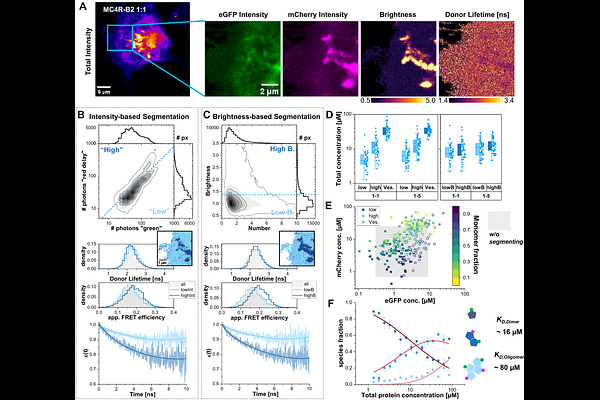
AbstractUnderstanding protein oligomerization in living cells is essential for elucidating cellular signaling and regulation, yet quantitative analysis remains challenging due to heterogeneous expression levels, dynamic interactions, and limited access to absolute protein concentrations. Here, we present a standardized, open-source framework for quantifying protein assemblies in living cells by integrating fluorescence lifetime and anisotropy imaging (heteroFRET and homoFRET) with molecular brightness-based concentration estimation and image analysis. Using natural variants of a vertebrate GPCR, the melanocortin-4 receptor (MC4R-A and MC4R-B2), as a model system, we demonstrate how to discriminate monomers, dimers, and higher-order oligomers, extract inter-fluorophore distance distributions, and determine association constants under physiologically relevant conditions in living cells. Standard fluorescent protein tags report on proximity and oligomerization via Homo- and HeteroFRET. Association constants are quantified using the variable protein expression in living cells and the spectroscopy readouts. By high-content imaging we overcome the biological noise and attain data qualities comparable to conventional biochemical in vitro assays. Intensity- and fluctuation-based segmentation further extends the accessible concentration range within individual cells, improving affinity analysis robustness. Our results establish quantitative image spectroscopy on living cells as quantitative tool for investigating protein-protein interactions under physiologically relevant conditions. All computational workflows are implemented in open-source software and are accompanied by detailed protocols and analysis scripts, enabling reproducible application and adaptation. Beyond GPCRs, this framework provides a practical and transferable methodology for quantitative studies on protein-protein interactions, mechanistic studies and drug discovery in complex cellular environments.