Evaluating the reliability of tools for mRNA annotation and IRES studies
Evaluating the reliability of tools for mRNA annotation and IRES studies
May, G. E.; Akirtava, C.; McManus, J.
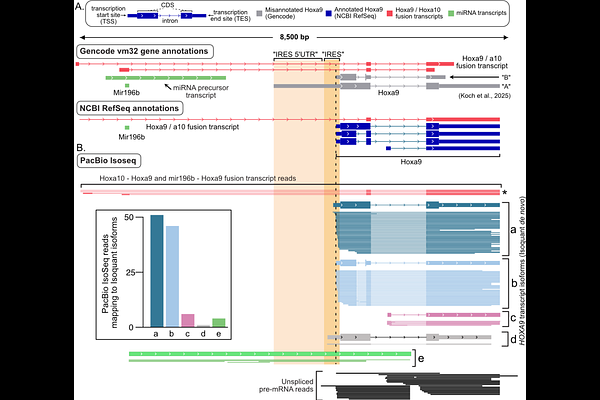
AbstractSince the discovery of viral Internal Ribosome Entry Sites (IRESes), researchers have sought to find similar elements in mammalian host genes, termed "cellular IRESes". However, the plasmid systems used to measure cellular IRES activity are vulnerable to false positives due to promoter activity in candidate IRESes. Orthogonal methods are needed to validate putative IRESes while carefully avoiding artifacts known to cause false positives. Recently, Koch et al. proposed approaches for studying IRESes, primarily circular RNA-generating plasmids, and for validating mRNA transcripts using smFISH and qRT-PCR. Here, we demonstrate confounding variables and artifacts in each of these approaches that can lead to inappropriate conclusions about potential cellular IRES activity. We show the back-splicing circRNA plasmid creates linear mRNA artifacts associated with false-positive IRES signals. Using orthogonal, gold-standard assays validated with viral IRESes, we find putative cellular IRESes reported using the back-splicing plasmid have no IRES activity. Furthermore, we demonstrate that smFISH and qRT-PCR can misidentify nuclear non-coding RNAs as mRNAs and we validate a single molecule sequencing assay for identifying genuine mRNA 5' ends. Our work establishes reliable methods for robust transcript annotation and IRES studies that avoid documented artifacts arising from bicistronic and back-splicing circRNA plasmid reporters.