A Modular Framework for Automated Segmentation and Analysis of AFM Imaging of Chromatin Organization
A Modular Framework for Automated Segmentation and Analysis of AFM Imaging of Chromatin Organization
Sorensen, E. W.; Pangeni, S.; Merino-Urteaga, R.; Murray, P. J.; Rudnizky, S.; Liao, T.-W.; Rashid, F.; Hwang, J.; Yamadi, M.; Feng, X. A.; Zähringer, J.; Gu, S.; Davidson, I. F.; Caccianini, L.; Osorio-Valeriano, M.; Farnung, L.; Vos, S.; Peters, J.-M.; Berger, J. M.; Wu, C.; Hatzakis, N. S.; Kirkegaard, J. B.; Ha, T.
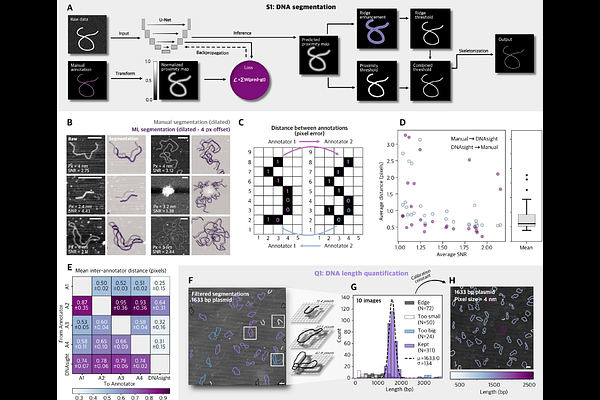
AbstractChromatin organization underlies essential genome functions, but its nanoscale organization remains challenging to capture and quantify with precision. Atomic force microscopy (AFM) offers direct structural readouts of DNA and chromatin, yet translating these rich images into reproducible biological metrics has been limited by the lack of standardized, scalable analysis tools. Here we present DNAsight, an automated analysis framework that integrates machine learning (ML)-based segmentation with modular, base-pair-calibrated quantification of DNA spatial organization, looping, nucleosome spacing, and protein clustering. Applied across diverse chromatin-associated proteins, DNAsight reveals protein-specific organizational signatures, including topology-dependent compaction by integration host factor (IHF), cofactor-mediated cohesin loop stabilization by precocious dissociation of sisters 5A (PDS5A), and promoter-driven multimerization of GAGA factor (GAF) clusters. The framework further enables direct extraction of nucleosome spacing distributions from raw AFM images, providing a label-free route to investigate chromatin fiber architecture. Together, these advances establish DNAsight as a generalizable and scalable approach for converting AFM measurements into quantitative insights into the physical principles of chromatin organization.