Genetic background shapes AI-predicted variant effects
Genetic background shapes AI-predicted variant effects
Schilder, B. M.; Liu, Z.; Desmarais, J. J.; Laub, D.; Rahimi, F.; Sethi, P.; Pereira, L. A.; Sun, M. M.; Kinney, J. B.; McCandlish, D. M.; Zhou, J.; Koo, P. K.
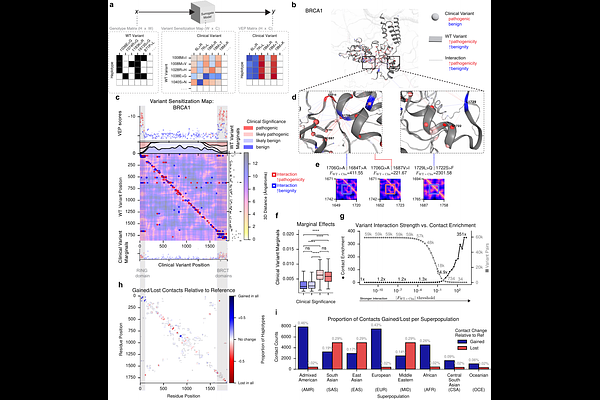
AbstractPredicting the consequences of genetic variants remains a major goal in biomedicine. Conventional approaches typically assess single-nucleotide variants in the context of a single reference genome, without accounting for genetic diversity that can modulate variant effects. Here we introduce the personalized variant effect predictor (pVEP) framework, which quantifies how genetic background across thousands of human genomes from globally diverse populations shapes computational predictions of clinical variant effects. Across deep learning models spanning protein structure, splicing, and noncoding regulation, pVEP reveals that many clinical variants exhibit heterogeneous predicted effects across haplotypes, with the same variant predicted to be pathogenic in some genetic backgrounds and benign in others. We find support for underlying molecular mechanisms, including shifts in predicted protein contacts and changes in splice-site recognition. Overall, personalized genomic context emerges as a systematically underappreciated variable in variant annotation and clinical interpretation, with particular implications for genetically diverse populations.